Section outline
-
-
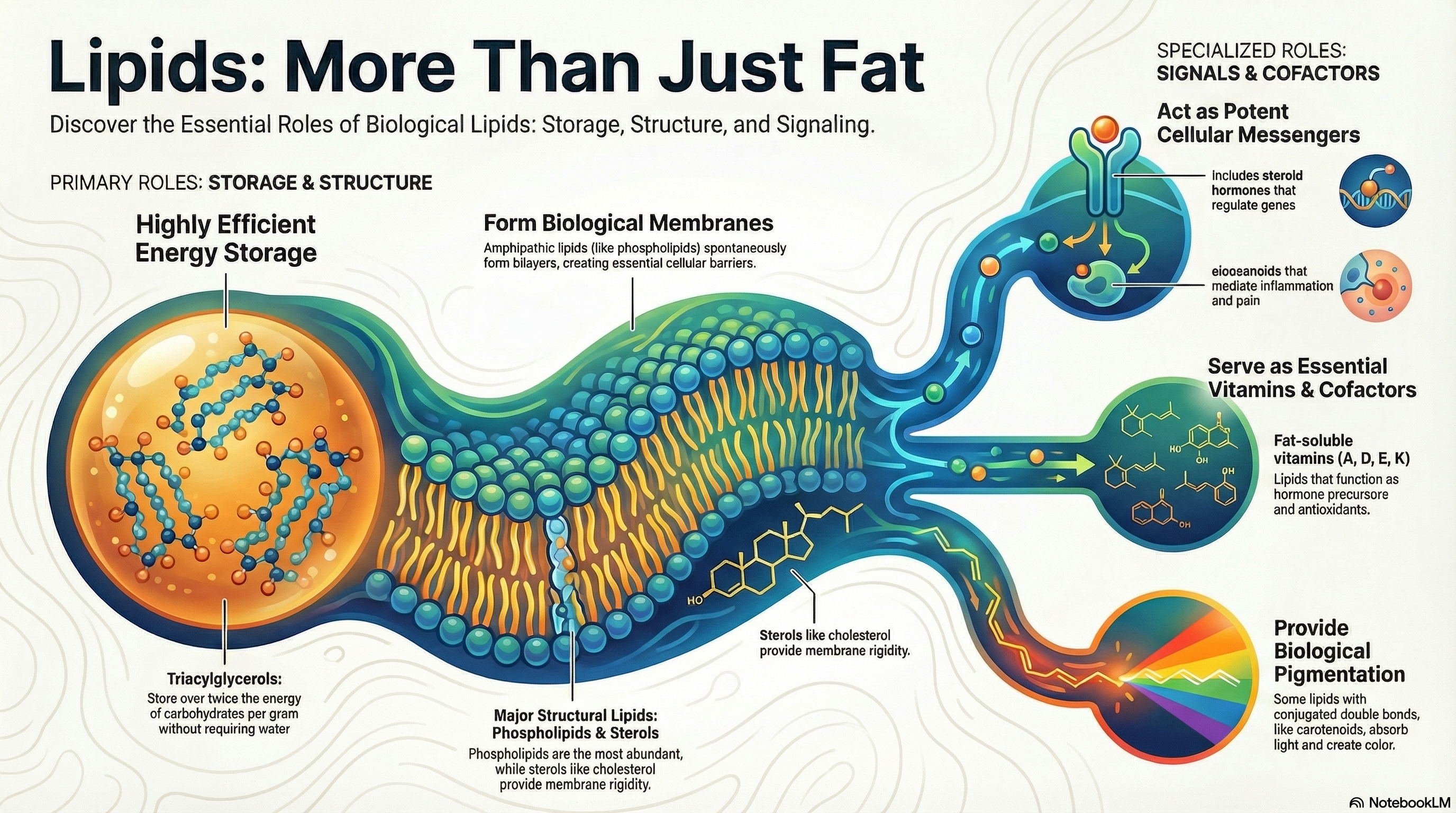
Lipids are a vast and chemically diverse group of biological molecules unified by a single defining feature: their insolubility in water. While this property may seem simple, it is the foundation for their critical and varied functions within living organisms. This guide provides an overview of the three primary biological roles that lipids fulfill. First, as fats and oils, they serve as the body's most efficient and compact form of long-term energy storage, packing more than twice the energy of carbohydrates per gram. Second, lipids are the fundamental architectural components of all cellular membranes. Their unique structure allows them to spontaneously form the lipid bilayer that encloses every cell, creating a vital barrier between the internal cellular environment and the outside world. Third, beyond these structural and storage roles, other lipids, though present in much smaller quantities, act as potent signaling molecules, hormones, and essential vitamins. These active lipids regulate a wide array of processes, from inflammation and blood pressure to gene expression and embryonic development. This document will explore each of these functions in detail, providing a clear and memorable foundation for understanding lipid biochemistry. -
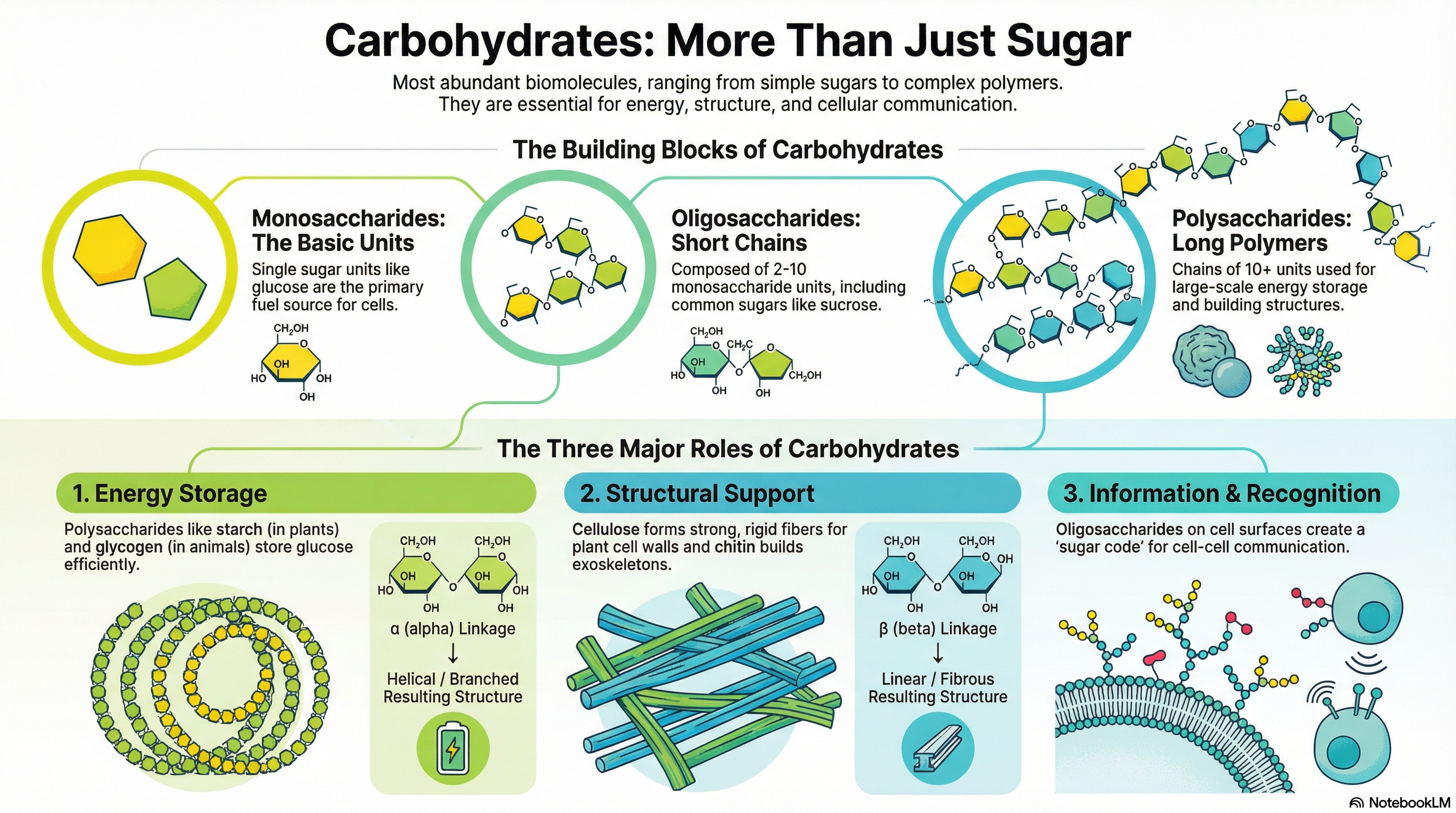
Carbohydrates are the most abundant biomolecules on Earth, serving as primary energy sources, structural components, and informational molecules in virtually all living organisms. Chemically defined as aldehydes or ketones with at least two hydroxyl groups, or substances that yield them upon hydrolysis, they range from simple sugars to complex polymers. The fundamental building blocks are monosaccharides, such as D-glucose, which can be linked together to form short chains called oligosaccharides (e.g., sucrose) or vast polymers known as polysaccharides.
The function of a polysaccharide is dictated entirely by its molecular architecture, specifically the type of monosaccharide units it contains and the nature of the chemical linkages, or glycosidic bonds, that connect them. This principle is best illustrated by comparing starch and cellulose. Both are polymers of D-glucose, yet they serve vastly different roles. Starch, linked by α(1→4) bonds, forms a compact helix that is easily digestible, making it an ideal energy storage molecule. In contrast, cellulose, linked by β(1→4) bonds, forms straight, rigid fibers that are indigestible by most animals, providing structural integrity to plant cell walls. This simple difference in stereochemistry—alpha versus beta—is a masterclass in biochemical principles, demonstrating how a subtle change in molecular geometry creates a profound difference in biological function: one stores energy, the other builds worlds.
-
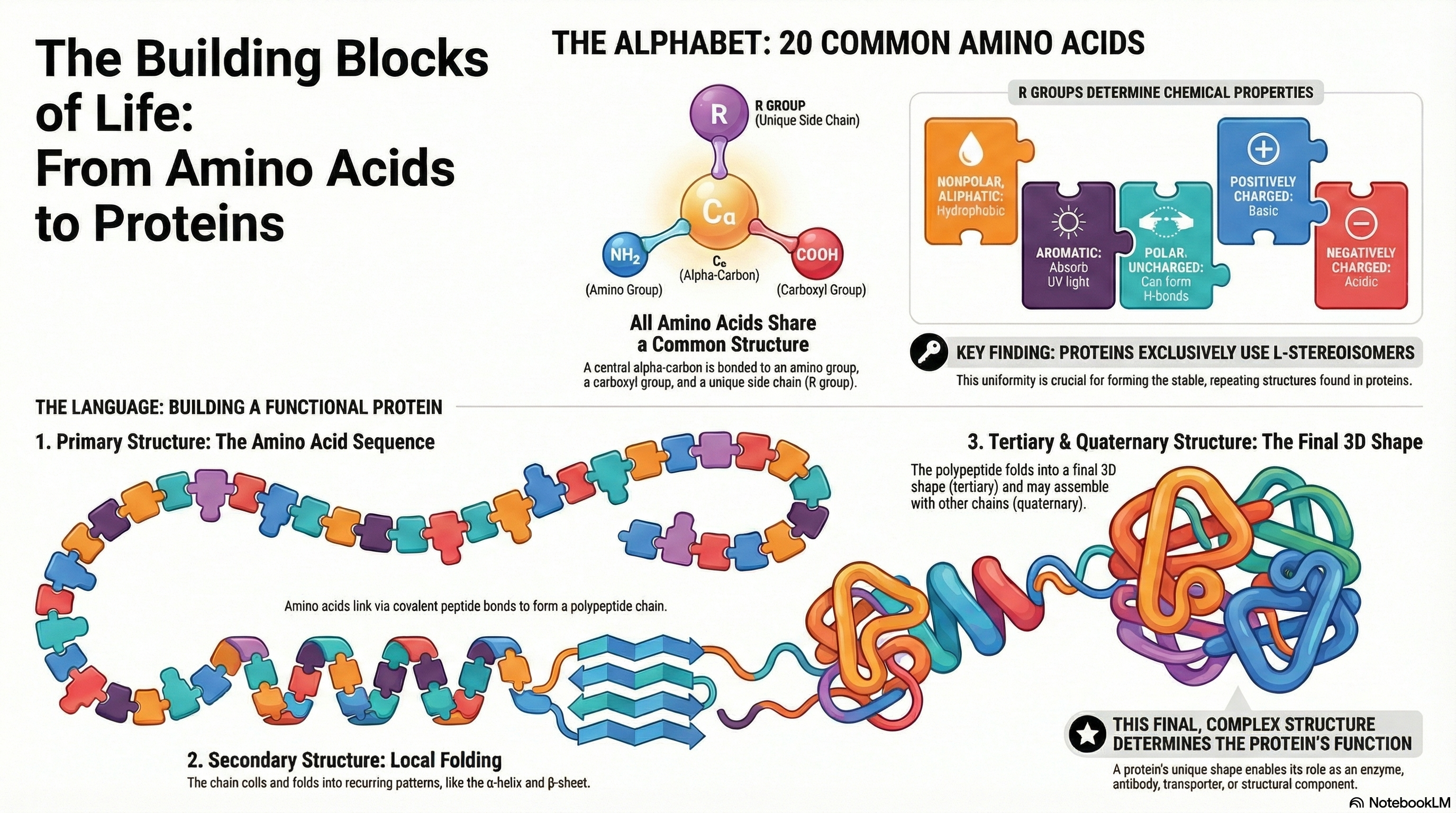
Proteins are the essential molecular machines of life, mediating virtually every process within a cell. Their incredible functional diversity, from catalyzing biochemical reactions to providing structural support, originates from a remarkably simple foundation: a common alphabet of 20 standard amino acids. Each amino acid shares a core structure but is distinguished by its unique side chain, or R group. The distinct chemical properties of these R groups—their size, shape, charge, and polarity—are the fundamental determinants of a protein's character.
These amino acid building blocks are covalently linked in a specific order via peptide bonds to form long chains called polypeptides. This defined sequence of amino acids constitutes the protein's primary structure. The primary structure is not merely a list of components; it is the blueprint that dictates the protein's entire architecture. Based on the chemical interactions between the amino acid side chains, the linear polypeptide folds into a specific three-dimensional structure, a complex architecture comprising secondary, tertiary, and in some cases, quaternary levels of organization.
Ultimately, it is this precise three-dimensional structure that endows a protein with its specific biological function. A single alteration in the amino acid sequence can disrupt this structure and abolish function, leading to disease. Therefore, understanding the foundational principles—the properties of the 20 amino acids, the nature of the peptide bond, and the critical relationship between sequence, structure, and function—is the key to comprehending virtually all biological processes at a molecular level.
-
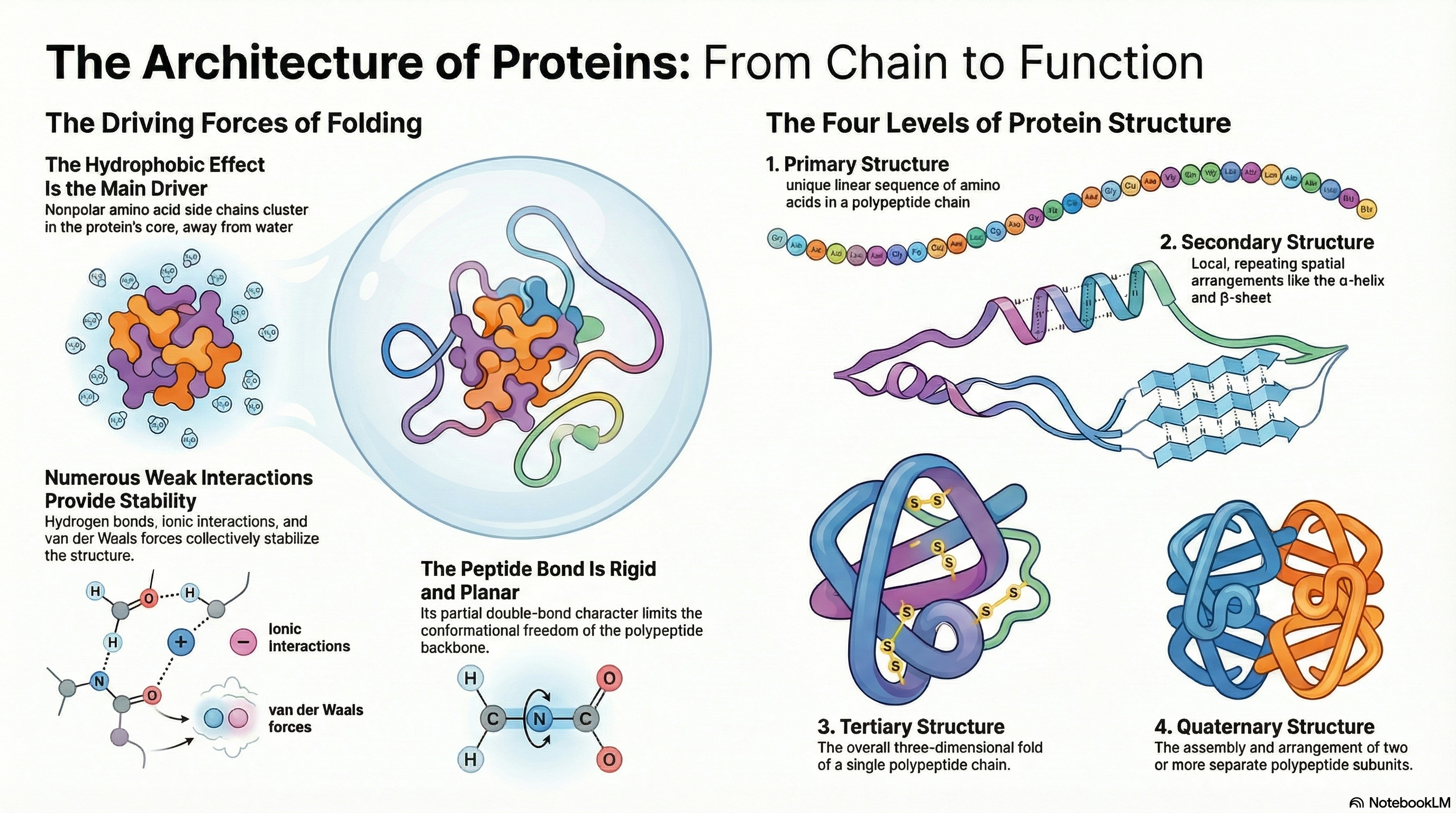
A protein's biological function is dictated by its precise three-dimensional architecture. This intricate structure is organized into a clear hierarchy. It begins with secondary structure, where local segments of the polypeptide chain coil into patterns like α-helices or arrange into β-sheets. These elements then fold into a specific, compact shape known as the tertiary structure, which defines the complete three-dimensional arrangement of a single polypeptide chain. For proteins composed of multiple chains, their assembly into a functional complex is described as the quaternary structure.
The stability of these structures relies not on powerful covalent bonds but on the cumulative effect of numerous weak, non-covalent interactions. While hydrogen bonds, ionic interactions, and van der Waals forces all play a role, the primary driving force behind protein folding is the hydrophobic effect. This is the thermodynamic tendency for nonpolar amino acid side chains to cluster in the protein's interior, shielded from water. This clustering releases ordered water molecules back into the bulk solvent, causing a favorable increase in the entropy of the water and making the folded state energetically favorable.
Remarkably, all the information required for a protein to achieve its final, functional 3D structure is encoded within its primary amino acid sequence. This fundamental principle was established by classic experiments showing that denatured proteins could spontaneously refold into their native, active state upon removal of the denaturing agent.
Correct protein folding is a matter of cellular life and death, as errors in this process have profound medical consequences. The misfolding and subsequent aggregation of proteins are the molecular basis for a wide range of debilitating human diseases, including Alzheimer disease, Parkinson disease, and the infectious prion diseases. This direct link between molecular structure and human health underscores the critical importance of understanding the principles that govern protein architecture.
-
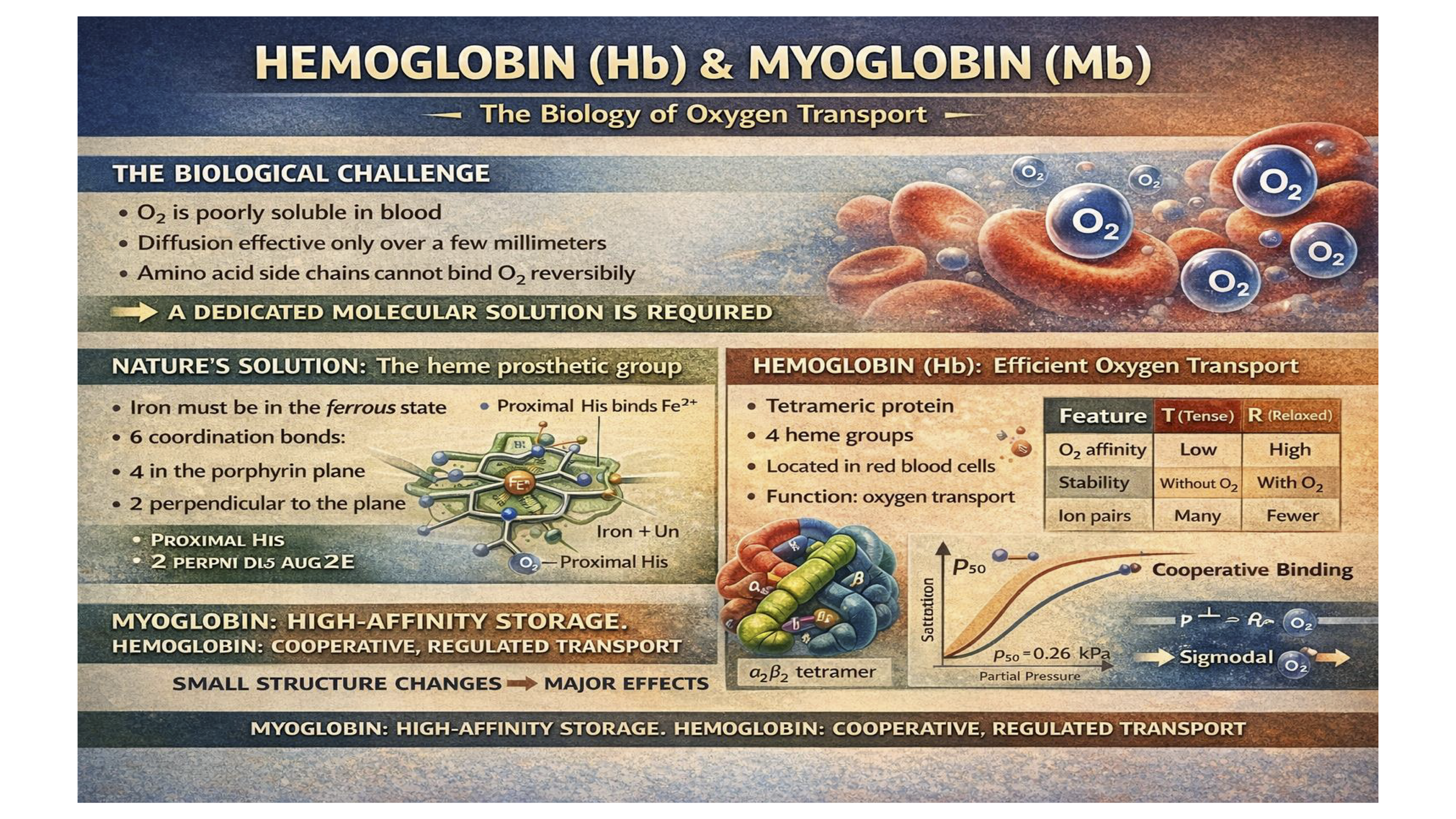
The central theme of this chapter is that proteins carry out their biological functions through dynamic and reversible interactions with other molecules, known as ligands. While some interactions are catalytic, this chapter focuses on non-catalytic binding, where the ligand's chemical composition remains unchanged. Myoglobin (Mb) and Hemoglobin (Hb) serve as the primary case studies for these principles. Myoglobin, a simple monomeric protein found in muscle, functions as an efficient oxygen-storage unit. Its high, constant affinity for oxygen ensures a ready supply during periods of high demand.
In contrast, hemoglobin is a complex, multi-subunit protein responsible for transporting oxygen in the blood from the lungs to the tissues. Its sophisticated function is a direct result of allosteric regulation. Hemoglobin can exist in two different structural states: a low-affinity "tense" (T) state and a high-affinity "relaxed" (R) state. The binding of one oxygen molecule to a subunit in the T state triggers a conformational change that is communicated to the other subunits, causing the entire protein to shift towards the R state. This process, known as cooperative binding, allows hemoglobin to bind oxygen efficiently in the high-oxygen environment of the lungs and release it effectively in the low-oxygen environment of the tissues.
The function of hemoglobin is further fine-tuned by several key physiological regulators, including pH, carbon dioxide (CO₂), and 2,3-bisphosphoglycerate (BPG). These molecules act as allosteric modulators, decreasing hemoglobin's oxygen affinity to promote its release where it is most needed. The take-home message is that the detailed three-dimensional structures of myoglobin and hemoglobin are perfectly adapted to their specific physiological roles. Even minor alterations to this structure, as demonstrated by the single amino acid change in sickle cell anemia, can have profound and devastating consequences on health.
-
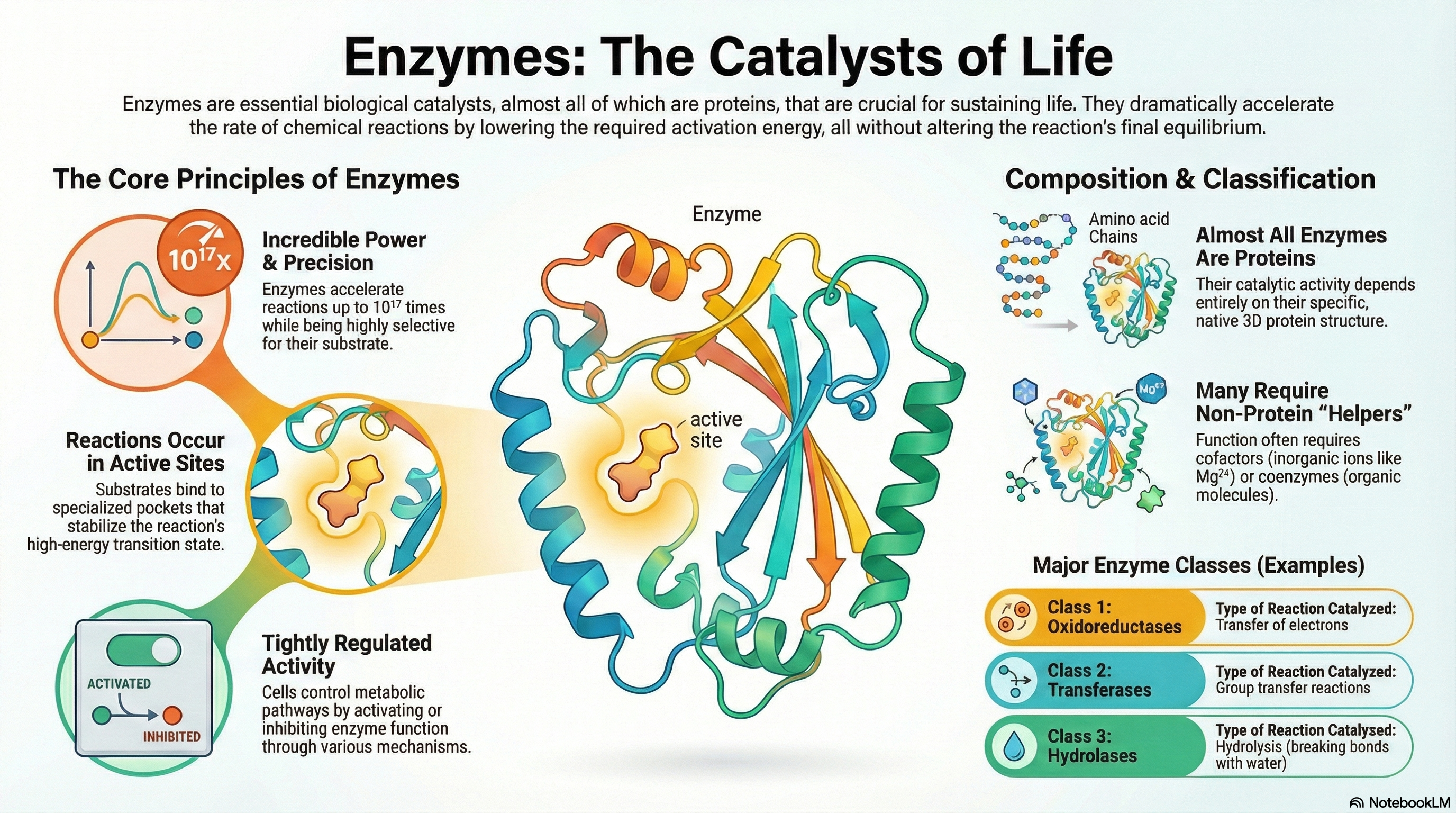
Enzymes are the essential biological catalysts responsible for accelerating the vast majority of chemical reactions necessary to sustain life. With the exception of a few catalytic RNA molecules, all known enzymes are proteins. Their function can be understood through five core principles. First, enzymes are extraordinarily powerful catalysts that dramatically increase reaction rates by lowering the activation energy barrier of a reaction; crucially, they do not alter the reaction's overall thermodynamic equilibrium. Second, they are highly specific, typically catalyzing only a single, specific chemical reaction or a few closely related ones. Third, these reactions take place within a specialized three-dimensional pocket on the enzyme known as the active site. Fourth, the immense catalytic power of enzymes derives from their ability to bind most tightly to the high-energy transition state of the reaction, effectively stabilizing it and facilitating the conversion of substrate to product. Finally, to ensure metabolic balance and responsiveness, the activity of enzymes within the cell is subject to tight and complex regulation.
Many enzymes cannot function alone and require non-protein components to perform their catalytic roles. These helpers are broadly classified as cofactors, which are inorganic metal ions (like Zn²⁺ or Mg²⁺), and coenzymes, which are more complex organic molecules that act as transient carriers of specific chemical groups. Many coenzymes are derived from essential dietary vitamins.
To bring order to the study of the thousands of known enzymes, a systematic classification system has been established. This system categorizes enzymes into seven major classes based on the type of reaction they catalyze, providing a logical and unambiguous framework for the field of enzymology.
-
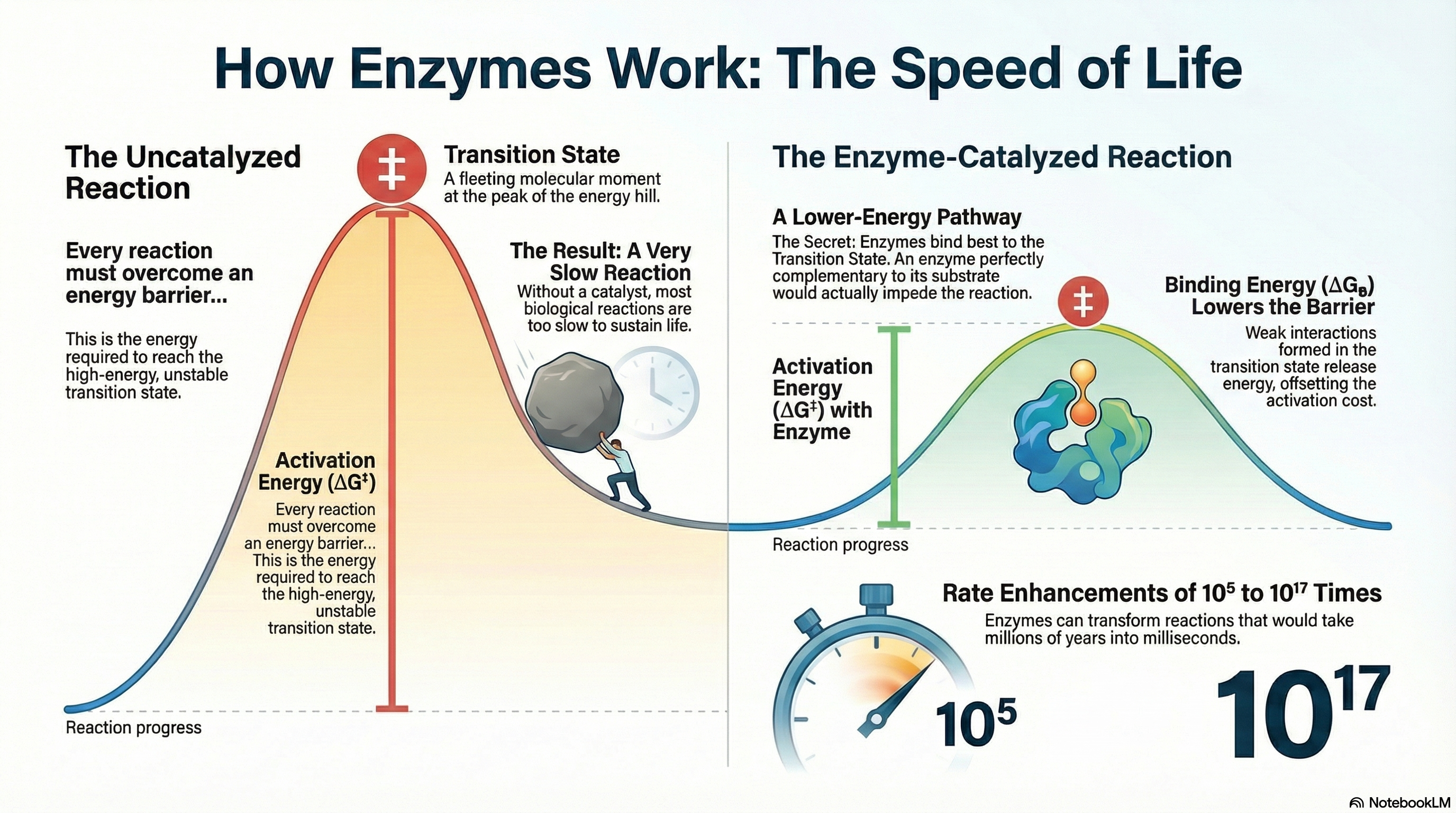
Enzymes are the essential biological catalysts that make life possible. While the molecules within cells are generally stable, the chemical reactions required for processes like digestion, energy production, and nerve signaling would occur far too slowly on their own to sustain an organism. Enzymes solve this problem by dramatically accelerating reaction rates by 5 to 17 orders of magnitude. They achieve this by lowering the activation energy (ΔG‡)—the energy barrier that must be overcome for a reaction to proceed. It is a fundamental principle of catalysis that while enzymes increase reaction rates, they do not alter the final reaction equilibrium. An enzyme helps a reaction reach its natural balance point much faster but does not change what that balance point is.
The primary source of an enzyme's catalytic power is binding energy (ΔGB). This is the free energy released when the enzyme forms multiple weak, noncovalent interactions (such as hydrogen bonds and ionic interactions) with the substrate. Crucially, enzymes have evolved not to bind the starting substrate molecule as tightly as possible, but rather to be structurally complementary to the high-energy transition state of the reaction. This means the strongest binding interactions are formed only when the substrate is distorted into this fleeting, unstable state. The favorable energy released from these interactions effectively "pays for" the energy required to reach the transition state, resulting in a much lower net activation energy and an extraordinary increase in reaction speed. To complement this powerful strategy, enzymes also employ specific chemical mechanisms, including general acid-base catalysis, covalent catalysis, and metal ion catalysis, to further facilitate the chemical transformation of the substrate.
VIDEO LINK -
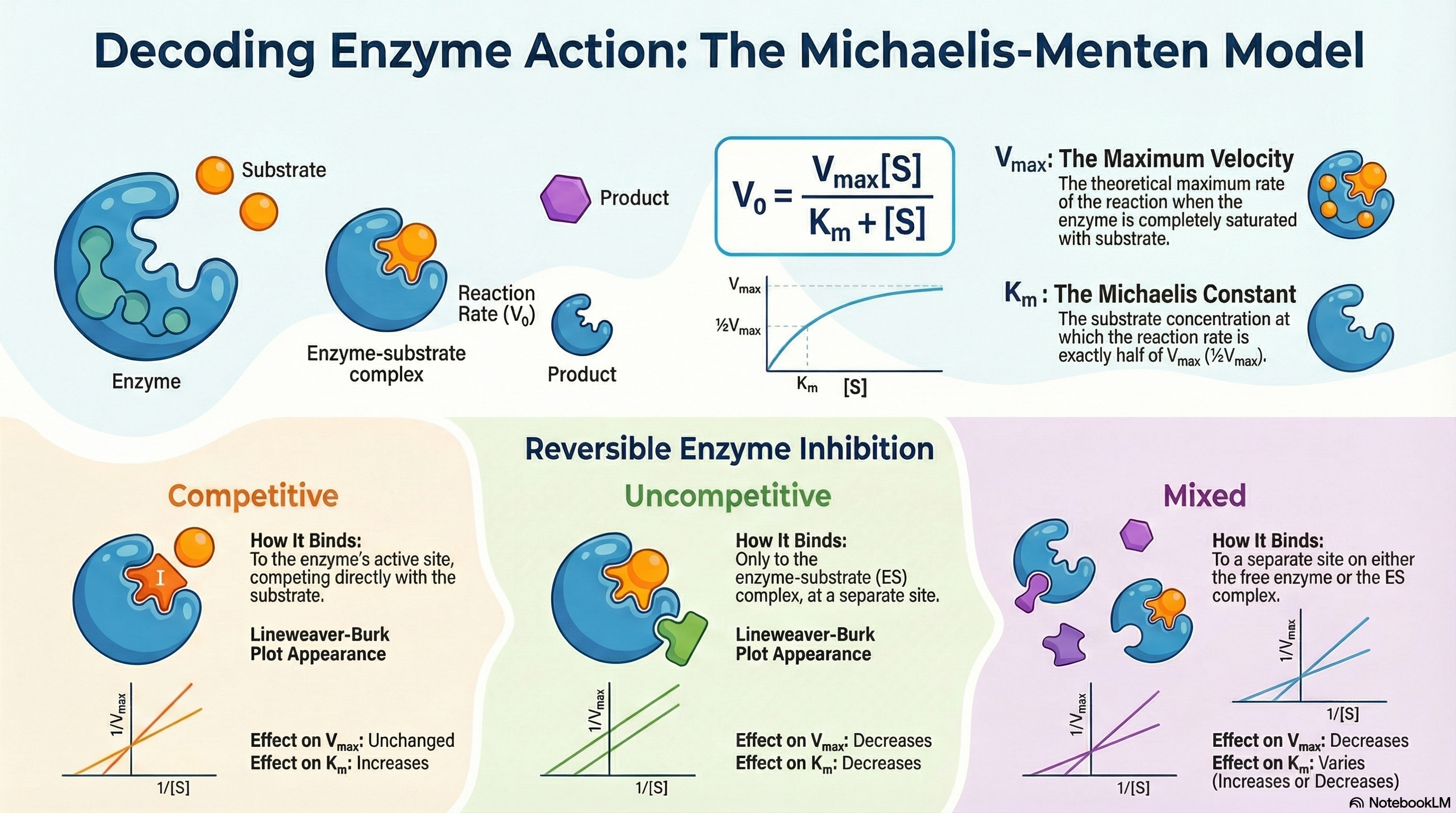
Enzyme kinetics is the quantitative study of the rates of enzyme-catalyzed reactions. This field provides critical insights into how enzymes function, complementing knowledge gained from structural and genetic analyses. At its core, the study of kinetics seeks to understand the relationship between the concentration of a substrate—the molecule an enzyme acts upon—and the speed, or velocity, of the reaction.
A key observation is that this relationship is not linear. As substrate concentration increases, the reaction velocity initially rises rapidly but eventually levels off, approaching a maximum velocity (Vmax). This plateau occurs because the enzyme becomes saturated; that is, all available enzyme molecules are bound to substrate and are working at their maximum capacity. Adding more substrate at this point cannot make the reaction go any faster.
The Michaelis-Menten model, proposed in 1913, provides the foundational mathematical description for this behavior. It is built on the concept of a two-step process: the rapid and reversible formation of an enzyme-substrate (ES) complex, followed by a slower, rate-limiting step where the complex breaks down to form the product and regenerate the free enzyme.
This relationship is quantified by the Michaelis-Menten equation, which uses two key parameters:
- Vmax (Maximum Velocity): The theoretical maximum rate of the reaction when the enzyme is fully saturated with substrate.
- Km (The Michaelis Constant): The substrate concentration at which the reaction proceeds at half of its maximum velocity (½Vmax). It is an indicator of an enzyme's efficiency at low substrate concentrations.
Furthermore, enzyme activity can be modulated by inhibitors—molecules that slow or stop catalysis. These are vital in medicine, as many drugs function as enzyme inhibitors. They are classified by how they interact with the enzyme and substrate, with distinct types like competitive, uncompetitive, and mixed inhibitors altering Vmax and Km in characteristic ways.
Ultimately, the Michaelis-Menten model remains a cornerstone of biochemistry. It provides an essential framework for characterizing enzyme performance, comparing catalytic efficiencies, and understanding the mechanisms of metabolic regulation and drug action.
-
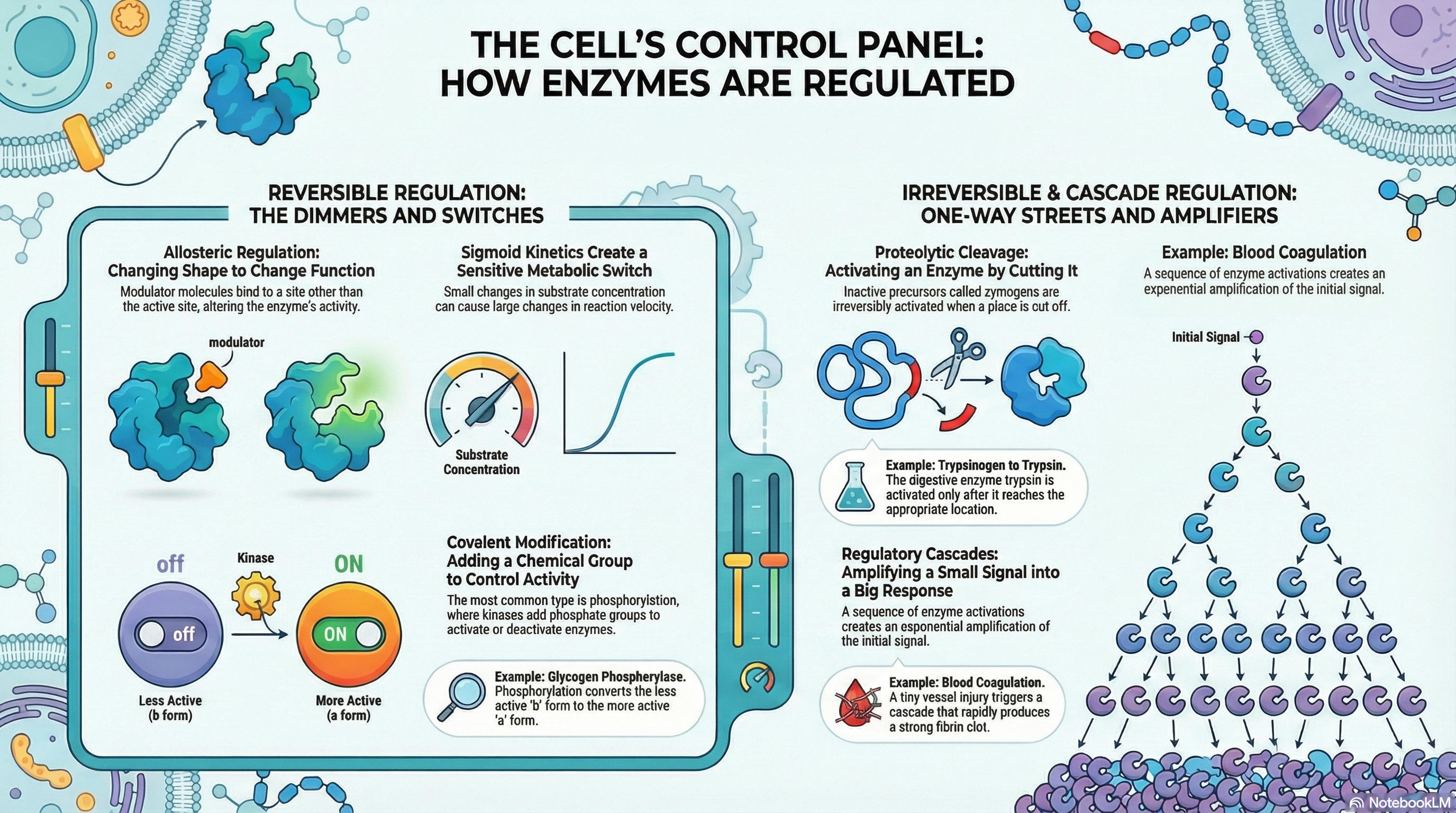
The persistence and adaptability of life depend on the precise control of countless biochemical reactions occurring within every cell. While enzymes are the catalysts that make these reactions possible, it is the regulation of their activity that orchestrates the complex symphony of metabolism. This document outlines the fundamental principles of enzyme regulation, explaining how cells dynamically adjust their internal chemistry to meet changing needs for energy, grow, and maintain a stable internal environment (homeostasis).
Cells employ four primary strategies to modulate enzyme activity. Allosteric regulation acts like a dimmer switch, where small molecules called modulators bind to a site on the enzyme separate from the active site, causing conformational changes that either increase or decrease its activity. Reversible covalent modification, most commonly the addition and removal of phosphoryl groups, functions as a rapid on/off switch, altering an enzyme's structure and function. Proteolytic cleavage offers an irreversible "on" switch, where inactive enzyme precursors, called zymogens, are permanently activated by the removal of a peptide segment; this is a common strategy for digestive enzymes and in the blood coagulation cascade. Finally, regulatory cascades create a powerful signal amplification system, where a small initial signal triggers a sequence of enzyme activations, leading to a massive and swift cellular response.
Crucially, these mechanisms are not mutually exclusive. Many key enzymes are subject to multiple layers of control, integrating signals from various metabolic pathways. This integration allows for exquisitely fine-tuned responses to complex cellular conditions. Understanding these regulatory mechanisms is fundamental to comprehending how cells manage resources, respond to their environment, and sustain the highly ordered processes we call life.
-
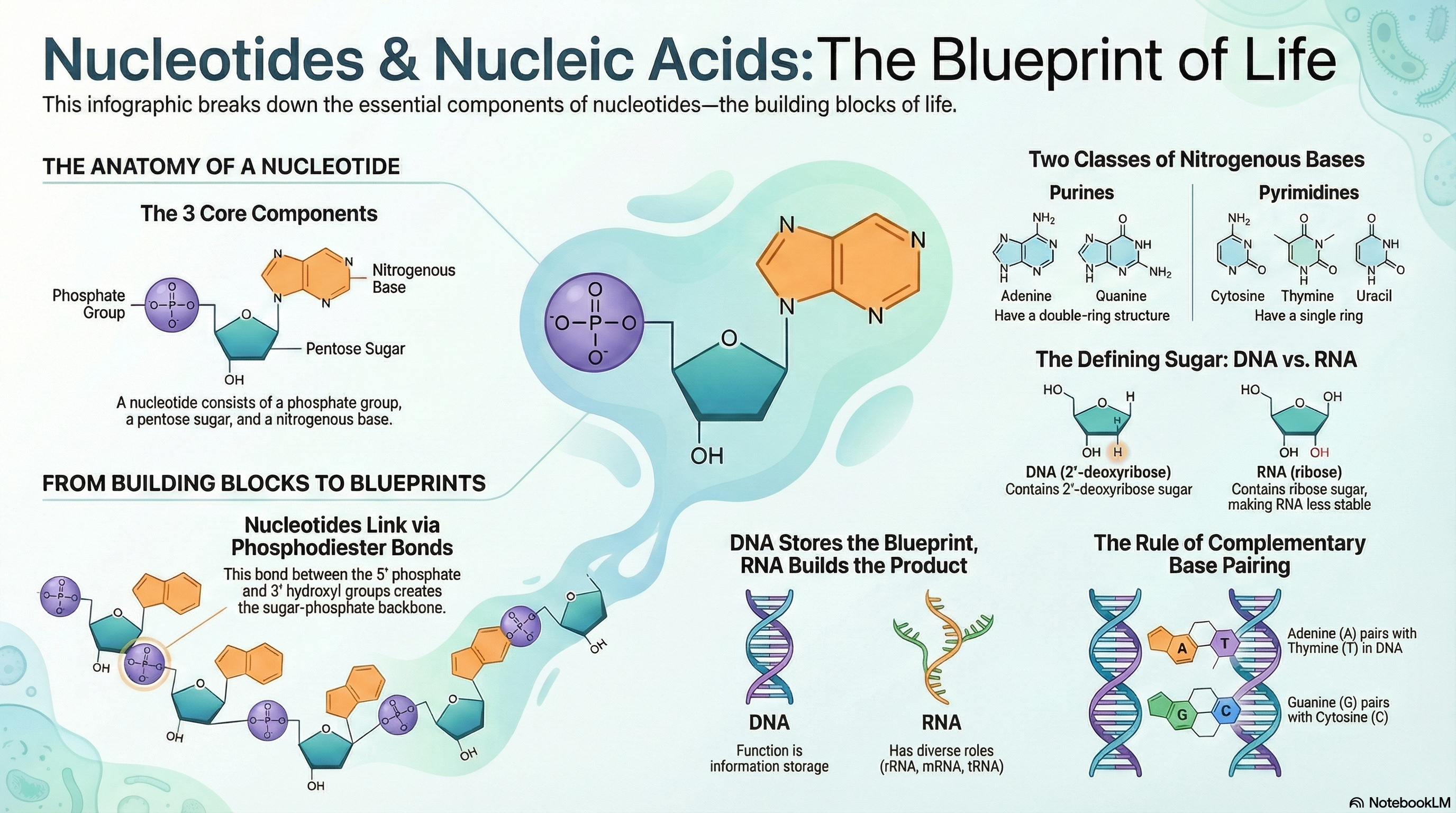
Nucleotides and their polymers, nucleic acids, are the cornerstones of life, serving a dual role as both the primary currency of cellular energy and the master molecules of biological information. At their core, these molecules are built from a simple three-part structure: a nitrogenous base, a five-carbon sugar, and one or more phosphate groups. While nucleotides like ATP power countless metabolic reactions, their most profound role is as the building blocks for deoxyribonucleic acid (DNA) and ribonucleic acid (RNA). The fundamental difference between these two information carriers lies in their sugar component—deoxyribose in DNA and ribose in RNA—a subtle chemical distinction that gives DNA its remarkable stability for long-term information storage and RNA its versatility and reactivity for more dynamic roles. The information itself is encoded in the sequence of the four bases within the polymer chain. The genius of this system lies in the principle of complementary base pairing, where Adenine (A) specifically pairs with Thymine (T) in DNA or Uracil (U) in RNA, and Guanine (G) always pairs with Cytosine (C). This precise, predictable pairing via hydrogen bonds is the molecular mechanism that allows genetic information to be copied with incredible fidelity, ensuring that the blueprint for life is passed accurately from one generation to the next.
-
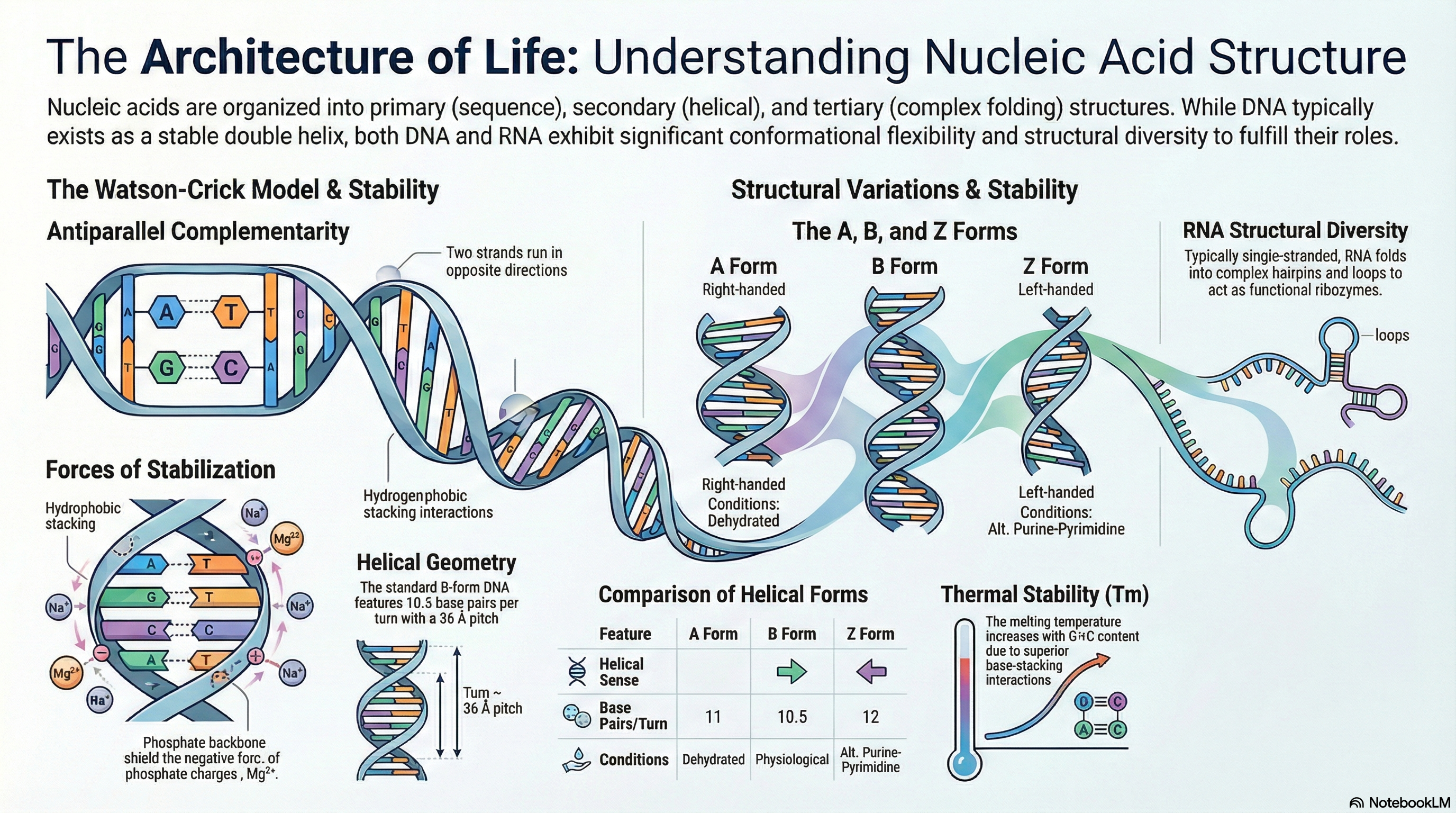
Nucleic acids, specifically deoxyribonucleic acid (DNA) and ribonucleic acid (RNA), are the fundamental molecules of life. They serve as the primary repositories and functional expressions of biological information, forming the blueprint that is transmitted from one generation to the next. DNA acts as the stable, long-term archive for genetic information, while RNA functions as the versatile messenger that translates this information into the functional machinery of the cell, such as proteins. Understanding the structure of these molecules is essential to comprehending how they perform these critical roles.
The following key principles provide a foundational overview of nucleic acid structure and stability:
- DNA's Structure: DNA is a right-handed double helix composed of two complementary strands that run in opposite directions (antiparallel). The molecule's hydrophilic sugar-phosphate backbones face outward, while the hydrophobic nitrogenous bases are stacked internally. This arrangement provides a stable framework for storing genetic information.
- Base Pairing Rule: The two DNA strands are held together by a precise rule of complementarity. The purine Adenine (A) always pairs with the pyrimidine Thymine (T) via two hydrogen bonds, and the purine Guanine (G) always pairs with the pyrimidine Cytosine (C) via three hydrogen bonds.
- Genetic Information: The specific sequence of these bases along a DNA strand constitutes the genetic code. The complementary nature of the double helix ensures that the information is stored reliably and provides a clear mechanism for its accurate replication, as each strand can serve as a template for synthesizing its partner.
- Structural Diversity: DNA is not a static molecule. While the Watson-Crick B-form is the most common under physiological conditions, DNA can adopt other conformations, such as the wider A-form or the left-handed Z-form, depending on hydration levels and nucleotide sequence. Furthermore, specific sequences can fold into unusual structures like hairpins, three-stranded triplexes, or four-stranded tetraplexes, which may have roles in gene regulation.
- RNA's Role and Structure: RNA primarily functions as a messenger (mRNA), carrying genetic instructions from DNA to the cell's protein-synthesis machinery. Although typically single-stranded, RNA can fold upon itself to create complex secondary structures, with the hairpin loop being the most common. This structural versatility allows RNA to perform a wide range of functions, including catalytic activity.
- Chemical Stability: The stability of the DNA double helix is a critical chemical property. The helix can be unwound, or "denatured," by heat or extreme pH. The temperature at which this occurs (the melting temperature, Tm) is largely determined by the molecule's Guanine and Cytosine (G+C) content. This is because G≡C base pairs exhibit stronger base-stacking interactions than A=T pairs, making G+C-rich DNA more stable.
VIDEO LINK
-
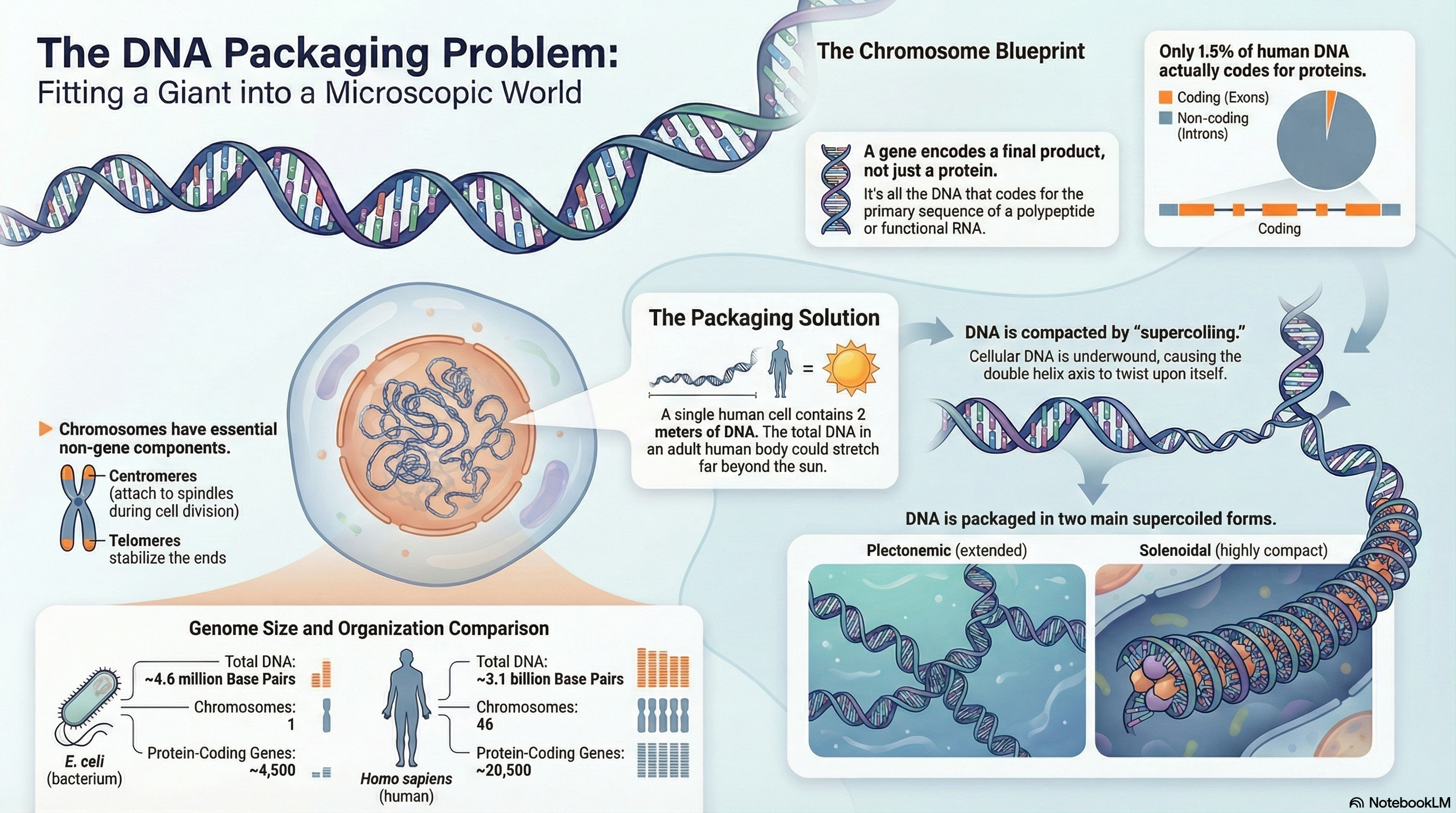
Every living organism faces a fundamental challenge of information management: how to package an immense length of DNA into a microscopic cellular space while keeping the genetic blueprint accessible for use. This summary explores the elegant, multi-layered solution to this problem, centered on the structure and organization of genes and chromosomes.
The modern biochemical definition of a gene is the entire stretch of DNA that codes for a final product, which can be either a protein or a functional RNA molecule. This genetic information is stored on DNA molecules that are orders of magnitude longer than the cells or viruses containing them. For instance, the DNA of a single human cell, if stretched out, would be two meters long. To solve this packaging problem, life has evolved sophisticated chromosomal structures. In prokaryotes like E. coli, the genome typically consists of a single, circular chromosome. In contrast, eukaryotes have much larger genomes divided among multiple linear chromosomes, which possess unique structural features. These include non-coding introns that interrupt the coding exons, a central centromere essential for cell division, and protective telomeres at the chromosome ends.
A key physical principle governing DNA compaction in all organisms is supercoiling. Cellular DNA is typically kept in an underwound state, meaning it has fewer helical turns than its most stable relaxed form. This creates torsional strain that causes the DNA to twist upon itself, forming supercoils. This "negative supercoiling" not only compacts the DNA but also stores energy that makes it easier for the two strands to separate—a critical step for DNA replication and transcription. In essence, chromosome structure is not a static storage unit but a dynamic, highly regulated system that masterfully balances the need for extreme compaction with the necessity of accessing genetic information to sustain life.
VIDEO LINK -
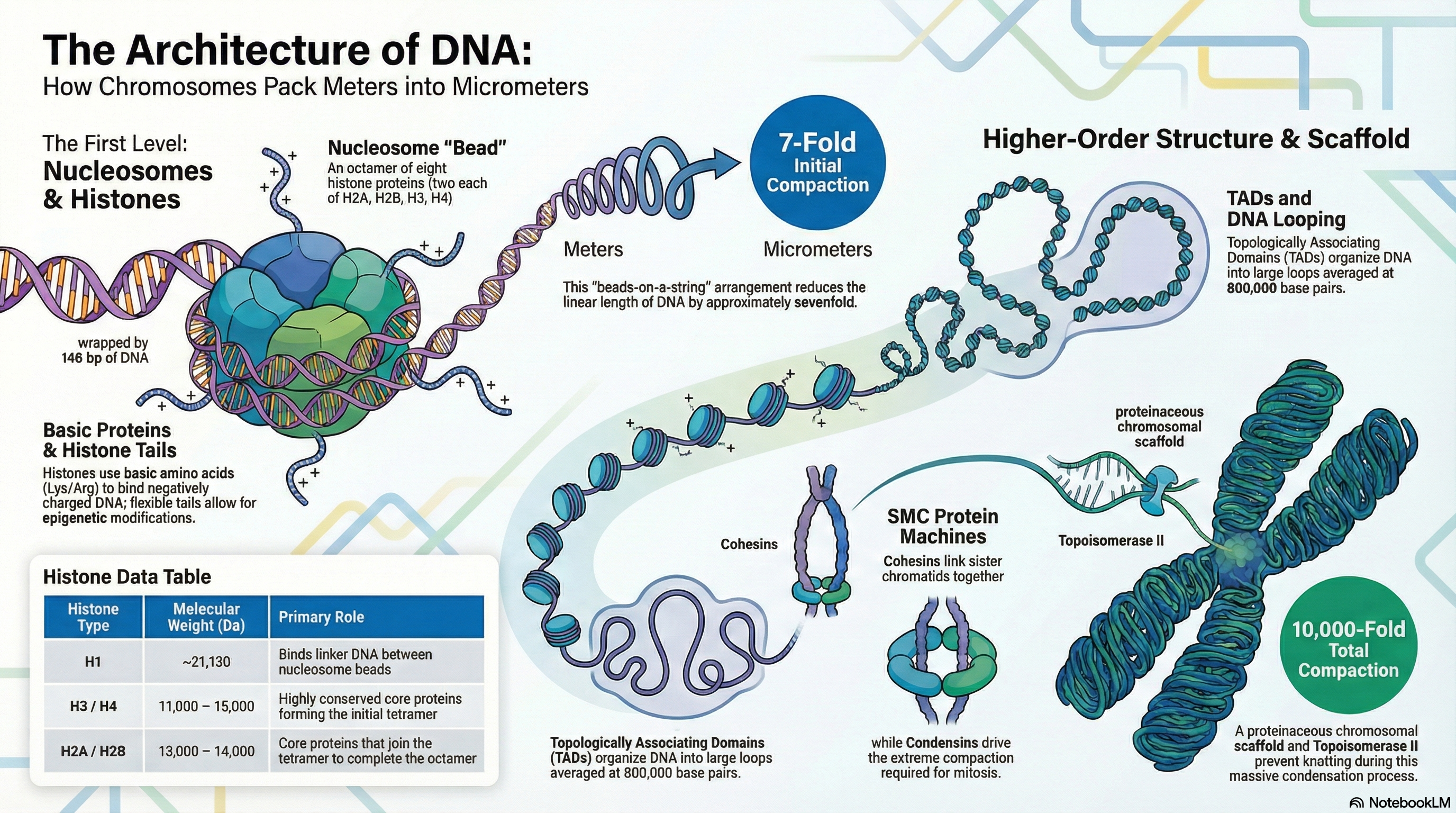
The packaging of genetic material presents a remarkable biological challenge: fitting several meters of DNA into a cell nucleus that is mere micrometers in diameter. This feat, equivalent to packing kilometers of thread into a small ball, requires a sophisticated and hierarchical system of organization. The fundamental material of eukaryotic chromosomes is chromatin, a complex of DNA, structural proteins, and RNA. The primary architects of this structure are a class of small, positively charged proteins called histones.
The first and most fundamental level of DNA compaction is the nucleosome, in which a segment of DNA wraps around a core of eight histone proteins, creating a structure often described as "beads-on-a-string." This initial step compacts the DNA approximately sevenfold. These nucleosomes are then folded into progressively more complex structures. Higher-order organization involves arranging the DNA into large loops, which are further segregated within the nucleus into functionally distinct compartments, such as transcriptionally active euchromatin and inactive, condensed heterochromatin. Ultimately, this multi-layered folding achieves a compaction of over 10,000-fold, culminating in the highly condensed chromosomes visible during cell division (mitosis).
Crucially, chromatin is not a static structure. Its organization is highly dynamic, constantly changing to accommodate the cell's needs. The structure decondenses to allow for DNA replication and gene expression during the cell's growth phase (interphase) and condenses dramatically for segregation during mitosis. This dynamic regulation, orchestrated by specialized proteins and RNA molecules, is essential for protecting the genome, ensuring its faithful inheritance, and controlling the flow of genetic information.
VIDEO LINK -
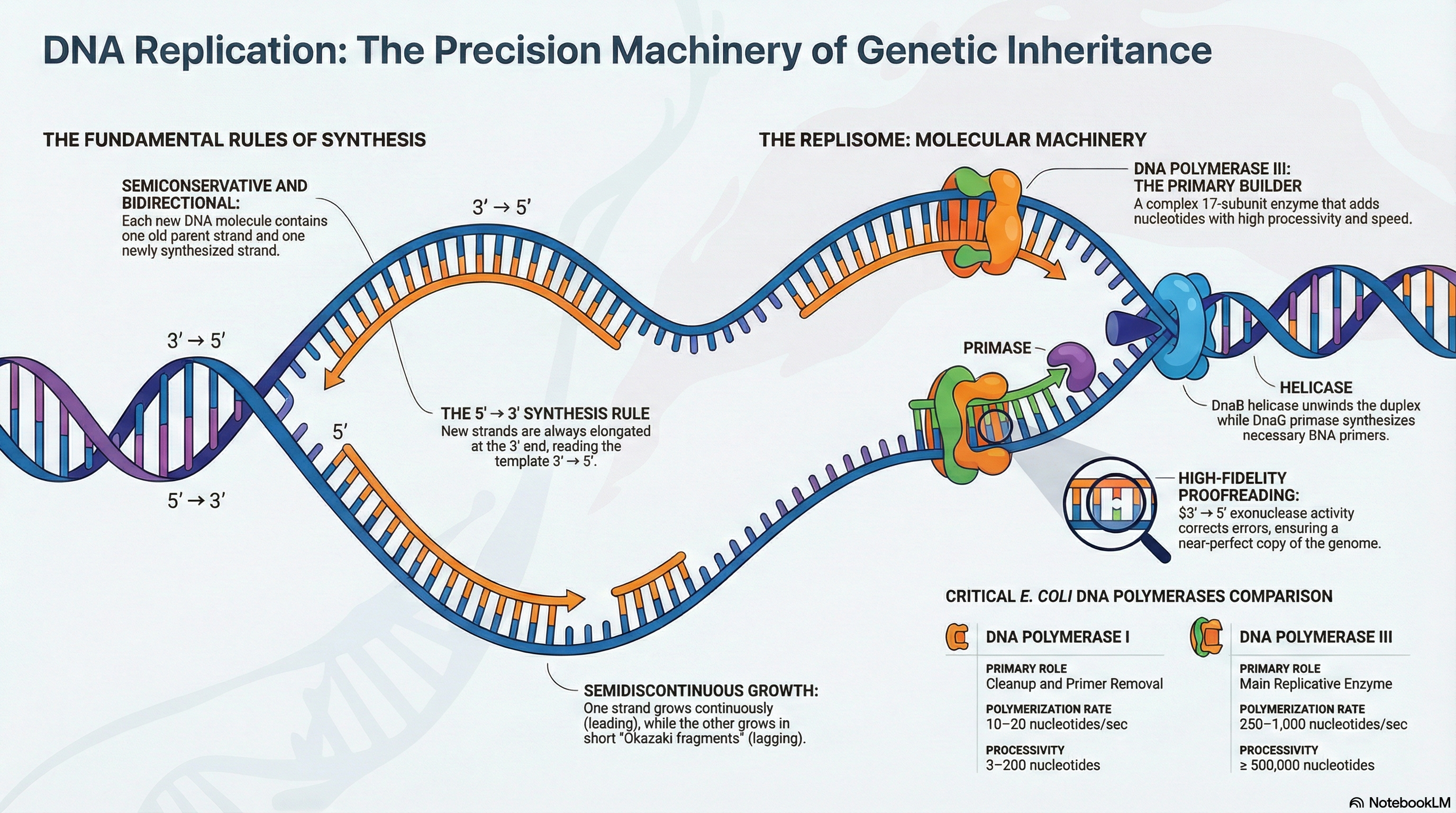
The integrity of an organism's genome is under constant threat from both internal and external forces, yet its faithful transmission from one generation to the next is a prerequisite for life. This chapter explores the suite of sophisticated molecular processes that cells have evolved to meet this fundamental challenge. The core of this system involves three highly integrated and essential pathways: DNA Replication, DNA Repair, and DNA Recombination.
DNA Replication is the process of accurately duplicating the entire genome before cell division. It operates under a strict set of rules, ensuring each new DNA molecule is a perfect copy of the original. This process is not simply about forming chemical bonds; it is about forming the correct bonds with extraordinary precision, a feat carried out by a complex machinery of specialized enzymes.
DNA Repair acts as the genome's quality control system. Despite the high fidelity of replication, errors occur, and DNA is constantly damaged by spontaneous chemical reactions and environmental factors. Cells employ a diverse array of repair pathways, each tailored to recognize and correct specific types of lesions. These systems are crucial for preventing the accumulation of mutations, which are strongly linked to diseases such as cancer. However, the rare errors that escape correction are not merely failures; they provide the raw genetic variation that serves as the fuel for evolution.
DNA Recombination involves the physical exchange of DNA segments between molecules. This process serves a dual purpose: it is a powerful mechanism for repairing severe DNA damage, such as double-strand breaks, and it is a primary driver of genetic diversity during meiosis in sexually reproducing organisms.
These three processes are not isolated events but are deeply interconnected aspects of DNA metabolism. This chapter focuses on the fundamental principles governing these pathways, the key enzymatic players that execute them, and their profound relevance to both evolution and medicine.
VIDEO LINK -
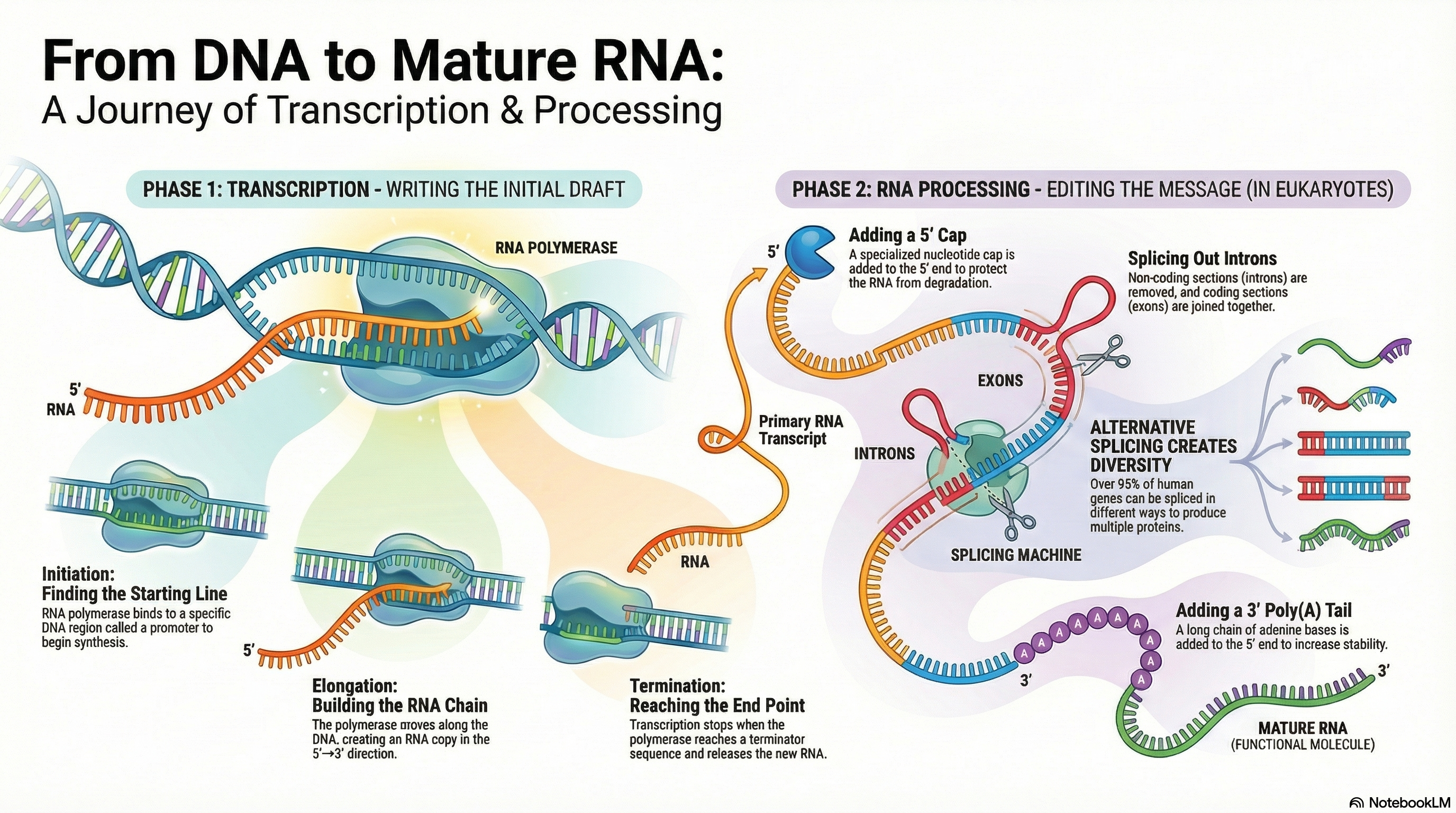
Ribonucleic acid (RNA) is a central molecule in the expression of genetic information, serving as far more than a simple messenger between DNA and protein. Its diverse roles as an information carrier, a biological catalyst, and a regulator of gene expression place it at the heart of cellular function. The primary process of RNA metabolism is transcription, the synthesis of an RNA molecule from a DNA template, catalyzed by the enzyme RNA polymerase. This process is highly selective, transcribing only specific segments of the genome into functional RNA molecules.
In eukaryotes, the initial RNA transcripts undergo a series of crucial modifications to become mature and functional. These post-transcriptional processing events include the addition of a protective 5' cap, the removal of non-coding sequences (splicing), and the attachment of a 3' poly(A) tail. These modifications are essential for the stability of messenger RNA (mRNA), its transport from the nucleus to the cytoplasm, and its efficient translation into protein.
The remarkable discovery that RNA itself can possess catalytic activity—acting as enzymes called ribozymes—revolutionized our understanding of molecular biology. This dual capacity to both store genetic information (like DNA) and catalyze chemical reactions (like proteins) provides the foundation for the "RNA World" hypothesis. This compelling theory posits that RNA, not DNA or protein, was the central macromolecule of early life, capable of storing a genetic blueprint while also catalyzing its own replication, thus solving the classic "chicken-and-egg" problem of which came first: the genetic blueprint or the functional machinery.
VIDEO LINK